Traditionally germs have been detected through microbiological methods, which are very precise but can take days or weeks. PCR- or antibody-based methods are rapid but require many steps and special equipment. “We were motivated to develop an especially simple, but very rapid and precise method,” says Li. “It must also be universal, meaning that it should be possible to develop tests for any desired germ using the same principle.”
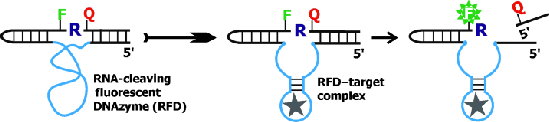 |
An RNA-cleaving fluorescent DNAzyme
(RFD) can produce a fluorescent signal in the crude
extracellular mixture generated by live bacterial cells
(see picture). These DNAzymes cleave a lone RNA linkage
(R) embedded in a DNA chain and flanked by nucleotides
labeled with a fluorophore (F) and a quencher (Q).
[Credit: Angewandte Chemie] |
“When a pathogen is metabolically active and multiplying in a given medium, it releases many substances into this environment. These are what we want to use,” says Li. The idea is to produce DNAzymes that react to a pathogen-specific product. A DNAzyme is a synthetic one-stranded DNA molecule with catalytic activity. Making a large pool of DNA molecules with random sequences and subjecting these to repeated selection and amplification steps allows for the development of molecules with the desired property.
At the core of the conceptual DNAzyme is a single RNA nucleotide. To its right and left are a fluorescing dye and a quencher. A quencher is a molecule that switches off the fluorescence of a dye when it is nearby. The researchers developed a DNAzyme that binds to a specific metabolic product from E. coli bacteria, which causes the DNAzyme to change its shape. In this altered form, the DNAzyme has RNA-splitting capability and cuts its own strand at the location of the RNA nucleotide. This separates the quencher from the dye, which begins to fluoresce. The fluorescence indicates that E. coli is present in the sample. This DNAzyme does not react to other bacteria.
“Through targeted selection, it should be possible to find a specific DNAzyme for any desired germ,” says Li. “It is not necessary to know what the metabolic product is, or to isolate it from the sample.” By using a common cell culture step, it is possible for the pathogens in a sample to multiply before the test, which allows for detection of a single original cell.



