Luminescent markers are an indispensable tool for researchers working with DNA. But the markers are troublesome. Some tend to destroy the function and structure of DNA when inserted. Others emit so little light, that they can barely be detected in the hereditary material. So researchers have been asking for alternative markers.
Now a PhD student at Department of Chemistry at the University of Copenhagen has developed a tool in collaboration with researchers at Chalmers Technical University, which could solve both problems: A tool that you might call a molecular gauge.
PhD student Soren Preus has investigated the properties of the two luminescent so called DNA base analogues tCO and tCnitro trying to determine whether they could measure the structure of DNA without disrupting it. His scrutiny has shown that the function of DNA is unimpeded by the insertion of the molecular gauge. And even better: One base analogue is very efficient at emitting light, and the other very good at receiving it. And because you can provoke transport of light-energy between the two luminescent markers they are usable for a measuring technique known as FRET or Fluorescence Resonance Energy Transfer.
Measuring angles with light
In brief FRET measurements are performed by forcing two luminescent markers to transfer light-energy from one to the other, and then measuring the efficiency of the transfer.
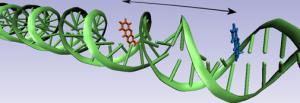
The two different markers are placed in the DNA-helix. When they are subjected to a lightpulse one marker (tCO) emits part of the energy to the other (tCnitro). This energy transfer can be measured. And by calculating backwards it is possible obtain very exact information about the distance and angle that the two have relative to one another.
Secrets from the center
Knowing distance and angle of the markers allows for calculations of distance and angle of all the natural base pairs in the DNA structure. And with that the researcher can put together a picture showing every twist and turn of the structure. Because structure and function are closely related in DNA, the method holds the potential to reveal new insights into the workings of DNA.
Putting the markers on the gauge
FRET-measurements are not a new phenomenon. What's new is, that Soren Preus has developed one of the base analogues tCnitro in collaboration with Swedish research institution Chalmers University of Technology. But even more important is the fact, that Mr Preus has used the facilities of the Molecular Engineering Group at University of Copenhagen to analyse every aspect of the energy-transfer between the two markers, because this allows future DNA-researchers to translate measurements to structure.
Fundamental as well as applied
Mr. Preus hopes that the new tool might find its use in characterising the structural changes that take place when a protein binds to DNA or RNA as that could explain basic cellular mechanisms. But equally important: The molecular gauge can be used to examine exactly how new drugs work, when they bind to DNA or RNA.
Further Information:
Soren Preus, Karl Brjesson, Kristine Kilsa, Bo Albinsson and L. Marcus Wilhelmsson:
Characterization of Nucleobase Analogue FRET Acceptor tCnitro.
In: Journal of Physical Chemistry B; J. Phys. Chem. B, 2010, 114 (2), pp 1050 - 1056, DOI 10.1021/jp909471b
Karl Brjesson, S. Preus, Afaf H. El-Sagheer, Tom Brown, Bo Albinsson and L. Marcus Wilhelmsson:
Nucleic Acid Base Analog FRET-Pair Facilitating Detailed Structural Measurements in Nucleic Acid Containing Systems.
In: Journal of the American Chemical Society; J. Am. Chem. Soc., 2009, 131 (12), pp 4288 - 4293, DOI 10.1021/ja806944w
Source: University of Copenhagen, Denmark
Last update: 08.04.2010
Perma link: https://www.internetchemistry.com/news/2010/apr10/fret-measurement-new-tool-developed-for-dna-research.php
More chemistry: index | chemicals | lab equipment | job vacancies | sitemap
Internetchemistry: home | about | contact | imprint | privacy
© 1996 - 2023 Internetchemistry
